ATCC Genome Portal Update: Gold Genomes, Tags, and Contig Information
2020 is off to an exciting start for the ATCC Genome Portal. One Codex and ATCC started the year on a strong foot after sequencing and closing 1 bacterial genome per day in 2019, and today we’re happy to announce three new features to help you leverage the data in the ATCC Genome Portal for your lab’s downstream processes.
Gold Genomes: Reference Genomes with Every Replicon Circularized
Closed and, where appropriate, circularized prokaryotic genomes are the gold standard of microbial genomics. These genomes are highly sought after for the complete context they offer, and they enable analyses such as massively improved prophage detection, comparative genomics of chromosomal and plasmid structure, and characterization of difficult-to-sequence repetitive regions.
ATCC’s hybrid approach to bacterial genome sequencing (using Illumina short reads for accuracy and Nanopore long reads for scaffolding, read more here) enables the assembly of these complete and appropriately circularized bacterial genomes wherever possible.
For those researchers looking to leverage the increased analytical power of closed bacterial reference genomes paired with ATCC standardized microbial materials, the ATCC Genome Portal now denotes these as Gold Genomes.
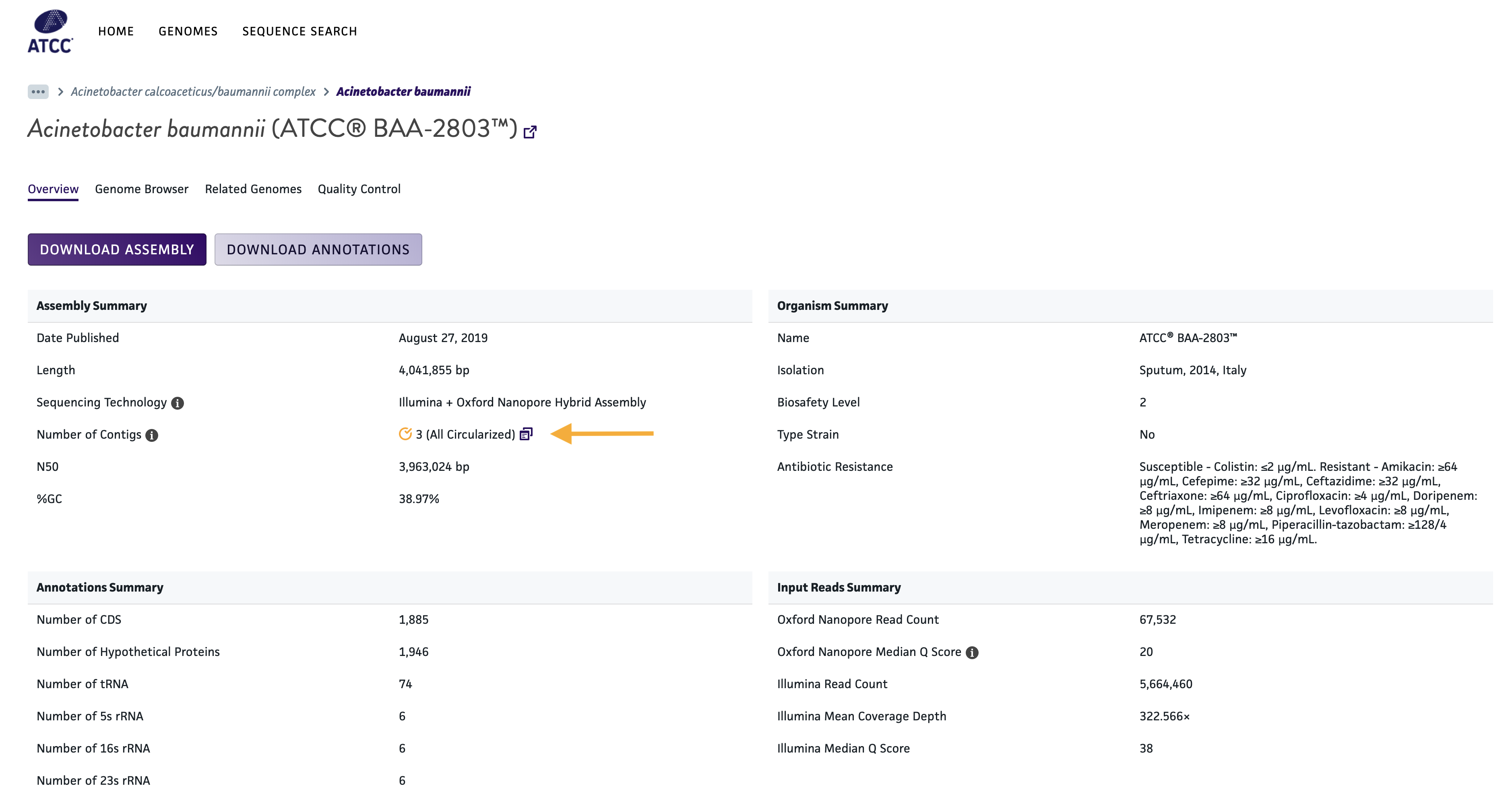
You can find Gold Genomes on the ATCC Genome Portal in two ways. First, if all contigs of a genome are complete and fully circularized, you’ll see the gold genome icon  displayed next to the contig count. Second, you can now add a new criterion to your genome portal search. Just add
displayed next to the contig count. Second, you can now add a new criterion to your genome portal search. Just add gold:yes to your search to only see Gold Genomes that match your query.
Tags: Explore ATCC’s Special Collections
ATCC’s bacterial collection is a treasure trove of unique and standardized microbial material. Contained within the main bacterial collection are groups of microbes that make up special collections; these are defined by their importance to prokaryotic nomenclature, inclusion in popular ATCC products, or their enhanced characterization.
To make it easier to see which genomes are part of these special collections, the ATCC Genome Portal now includes tags next to these genomes. Currently you’ll be able to find tags for three special collections:
1. Type Strains: genomes corresponding to strains with official standing in prokaryotic nomenclature, essentially the “definition” of the species. 
2. MSA Component: genomes of ATCC strains that are part of the popular ATCC® Microbiome Standard products. 
3. GPS: genomes of ATCC strains that are part of the “Global Priority Superbug” collection; these AMR-relevant strains include enhanced characterization data such as antibiotic susceptibility profiles and MIC values. 
To date, we’ve sequenced 27 Global Priority Superbug genomes, 29 MSA Component genomes, and 200 Type Strain genomes.
Just like with Gold Genomes, you can include these tags in your search; for example, simply add Tag:‘Type Strain’ to your search to only see Type Strain genomes that match your query.
Contig Tables: Rapidly Access Contig-Level Data
As we continue to develop the ATCC Genome Portal, we’re constantly listening to feedback to make the site more user friendly. One frequent request was a fast method to get information about the contigs that make up our genome assemblies.
We’re happy to make available a new, quickly accessible contig table, which can be found in the Overview tab of each genome on the portal. Simply click the icon next to the number of contigs listed for a particular genome to bring up a downloadable table, featuring contig names, their individual sizes, and whether the contig is circular or linear.
The ATCC Genome Portal: Ever Growing, Always Improving
Start using these new features and our ever-expanding list of high-quality reference genomes paired with ATCC standardized material today by signing up for an account or logging in with your existing One Codex account.
If you want to share feedback about the ATCC Genome Portal, request a specific feature, or request a genome for your favorite ATCC strain, we’d love to hear from you. You can find a feedback survey here.
If you have any questions about the ATCC Genome Portal, or need help navigating the database, ATCC Support is ready to help. Just fill out some quick details about your question here, and a representative will get back to you shortly.