Launching out of beta!
Today is a big day for One Codex, where we’re launching out of beta!
This marks the introduction of a number of new features (and many under-the-hood improvements), and incorporates much of the feedback our users have provided this year. You will notice some changes to our website, and the platform can now be found at app.onecodex.com (note that all previous links to the beta site will still work).
In addition to a number of improvements to the underlying data infrastructure of the platform, we are continuing to add new features that we think will increase your ability to generate useful insight from microbial genomic data.
Robust data infrastructure and HIPAA-level security
The One Codex platform now offers HIPAA-level security, with all genomic data and metadata now encrypted in transit and at rest. With these changes come the ability to safely handle regulated data (e.g., protected health information) as well as a broader set of security guarantees. If you would like to work with clinical data, please get in touch so we can discuss a HIPAA-compliant setup. With audit trails and highly reproducible analyses, One Codex is an ideal platform for clinical microbiology and metagenomics.
Expanded analysis with in silico Panels
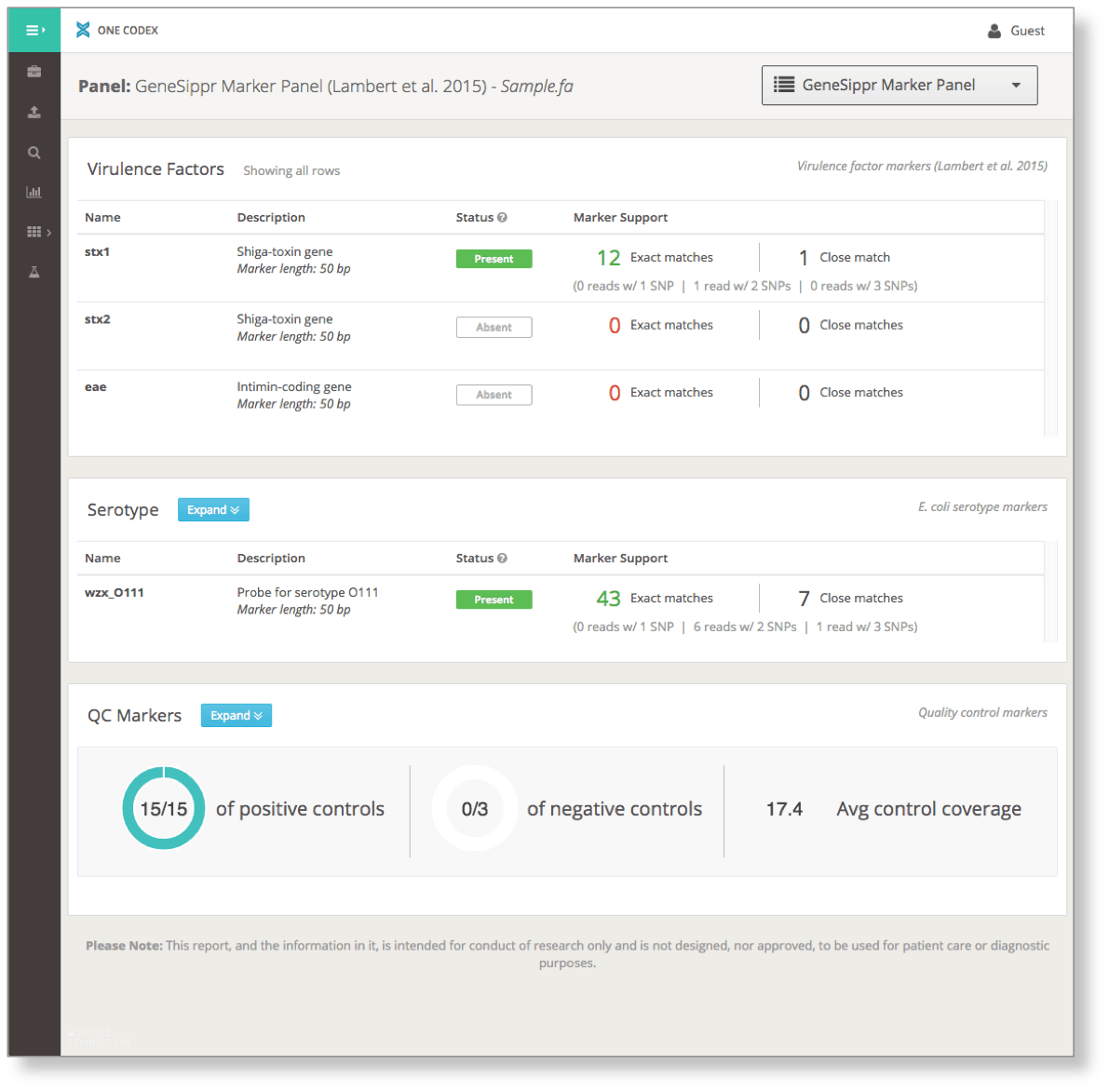
In silico gene or marker panels offer a functional window into an NGS sample, including predicted virulence factors, antibiotic resistance, and other phenotypic traits. This expanded analysis complements One Codex metagenomic analysis with specific, easy-to-interpret assays.
For example, this in silico panel uses a set of markers described by Lambert, et al. (2015) to characterize the virulence and serotype of Shiga toxin-producing E. coli (STEC), summarizing the results in a single comprehensive display:
Tag, organize, and search!
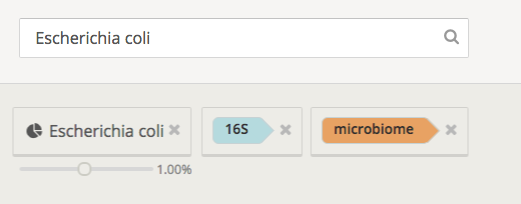
Finally, you can add tags to your datasets (we add some automatically, like “isolate” and “complex”), and use them to search and organize. You can also search through and filter your datasets by name, tags, and the organisms (or taxa) present. For example, the below search returns all datasets with the “16S” and “microbiome” tags and >1% E. coli present:
There are many more exciting things in the works here at One Codex, but we are also proud of where the platform stands today. We absolutely could not have built this platform without the valuable input of all our early users, and you have our thanks.
There are many more exciting things in the works here at One Codex, but we are also proud of where the platform stands today. We absolutely could not have built this platform without the valuable input of all our early users, and you have our thanks.
We are looking forward to exploring the future of microbial genomics with all of you, and as always, please don’t hesitate to be in touch.