Instant sequence search
We’re pleased to announce that we’re launching a new feature on One Codex today: instant sequence search.
With this new tool, simply enter a nucleotide sequence – whether a read from a FASTQ file or a section of a reference genome – and we’ll instantly compare it against our One Codex database.
How it works
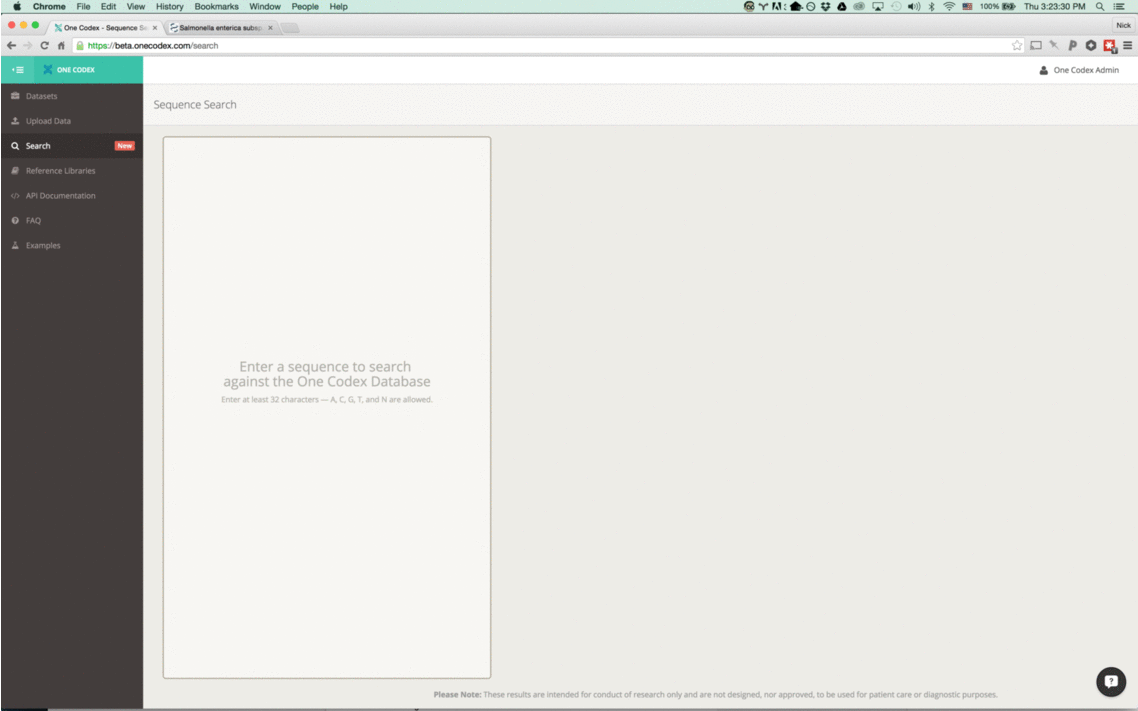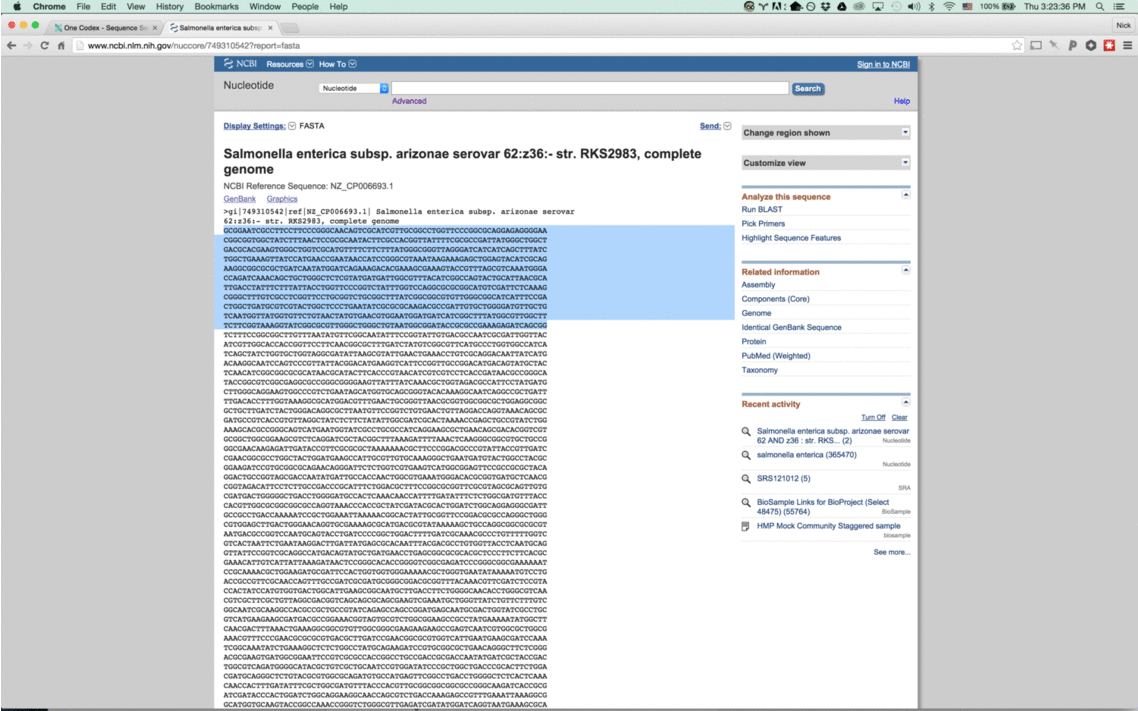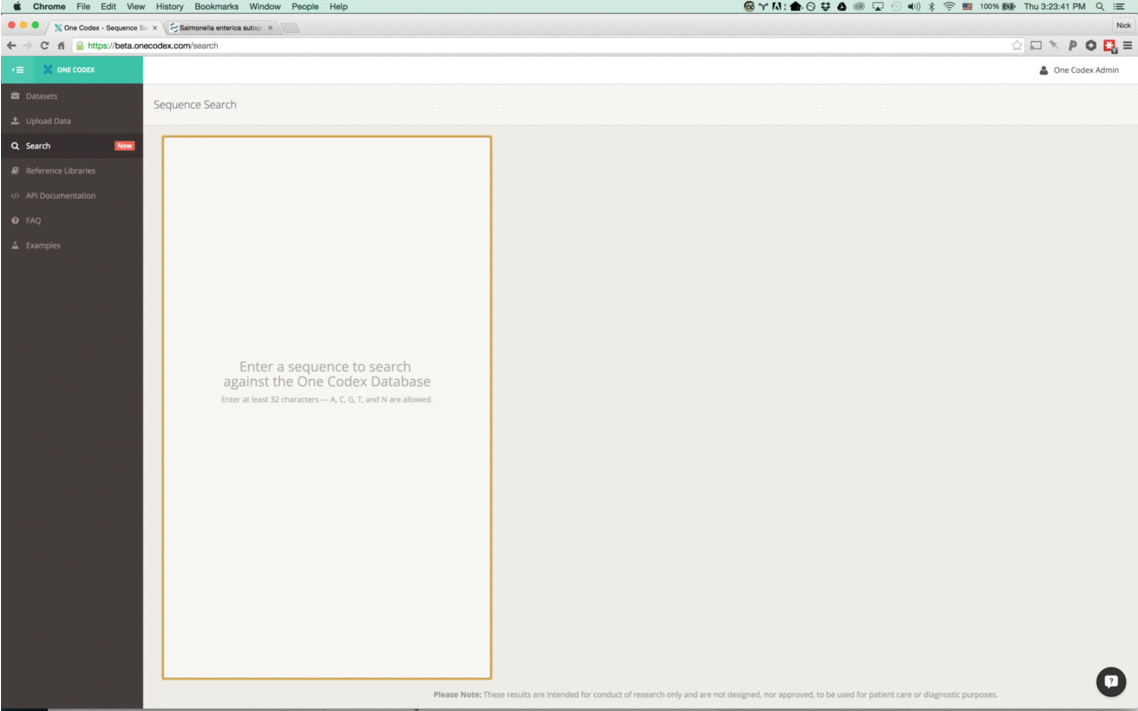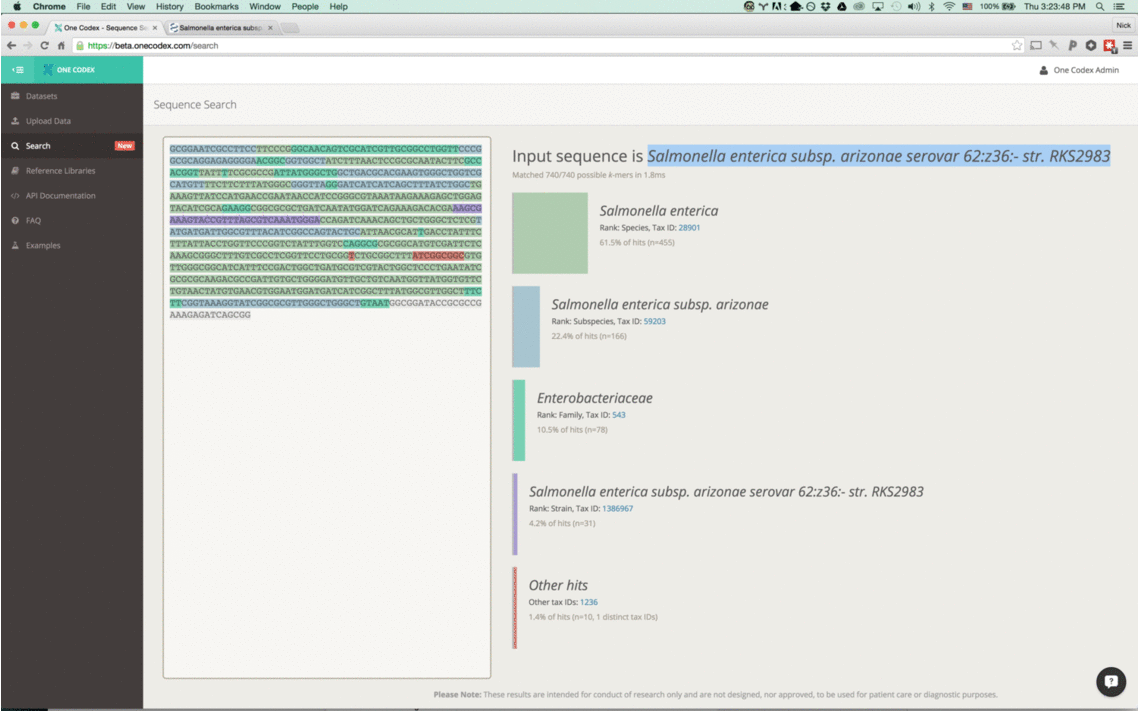
Our instant classifier works by splitting up the input sequence into all overlapping k-mers and then comparing each against our reference database, using a k=31. We then visually highlight the most specific taxonomic value (or lowest common ancestor) for the k-mer in our database. (Note that because the k-mers overlap, we only color the first character of each)
These results are then summarized on the right of the search display, including our classification of the overall sequence. This is Salmonella enterica subsp. arizonae serovar for the example above.
Your feedback & questions
We’d love to hear if you find this functionality useful, whether you’d like to see it in our API, and any questions you may have. As always, please feel free to drop us a note, follow us on Twitter, or chat with us directly on the site.