CDC "No-Petri-Dish" Challenge
We’re excited to announce that the CDC’s Advanced Molecular Detection initiative has named One Codex as the winner of its $200K “No-Petri-Dish” Challenge! We are thrilled to be recognized by a world-leading public health agency and excited to begin offering solutions for next-generation public health and epidemiology.
The “No-Petri-Dish” Diagnostic Test Challenge asked participants to develop a fast “straight-to-strain” method for identifying a type of E. coli known as STEC (a common cause of severe foodborne outbreaks). Current tests available to public health officials fall into two major categories: “culture-based” tests, which require growing a bacterial culture and can take days or weeks to complete; and faster, but significantly less informative tests for specific pathogens or toxins (e.g., PCR-based tests).
Our approach uses next-generation sequencing (NGS) data to offer fast, high-resolution diagnostic results. We developed a unique solution that is effective even for metagenomic samples with extremely low concentrations of E. coli or other pathogens (<= 1X coverage). This ultimately promises to help public health officials identify, track, and monitor outbreaks in near real-time.
This work for the CDC builds on the One Codex beta platform, where we’re building the largest index of genomic data and better search algorithms for fast, accurate bacterial and viral identification. We plan to add this functionality to our live beta soon, which we’ll announce here alongside some more technical details on our approach.
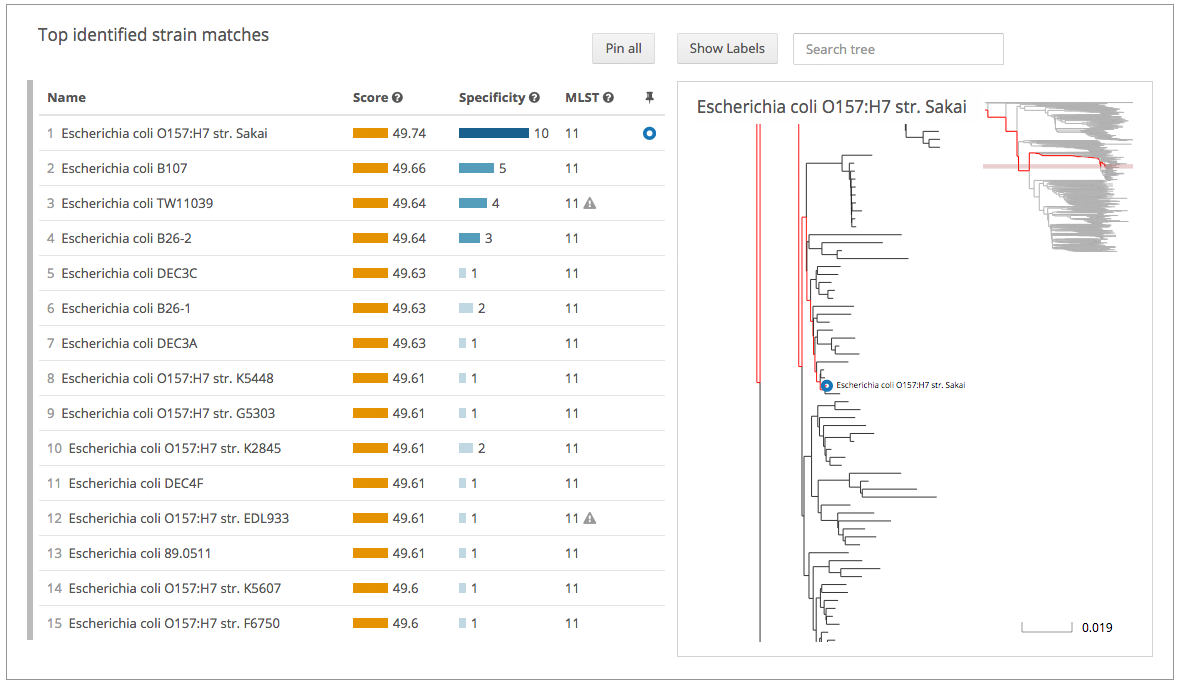 A strain typing example on the One Codex beta platform
A strain typing example on the One Codex beta platform
Ps. Please feel free to drop us a note if you’re working in the space – we’re always excited to connect with others working on sequencing-backed diagnostics.